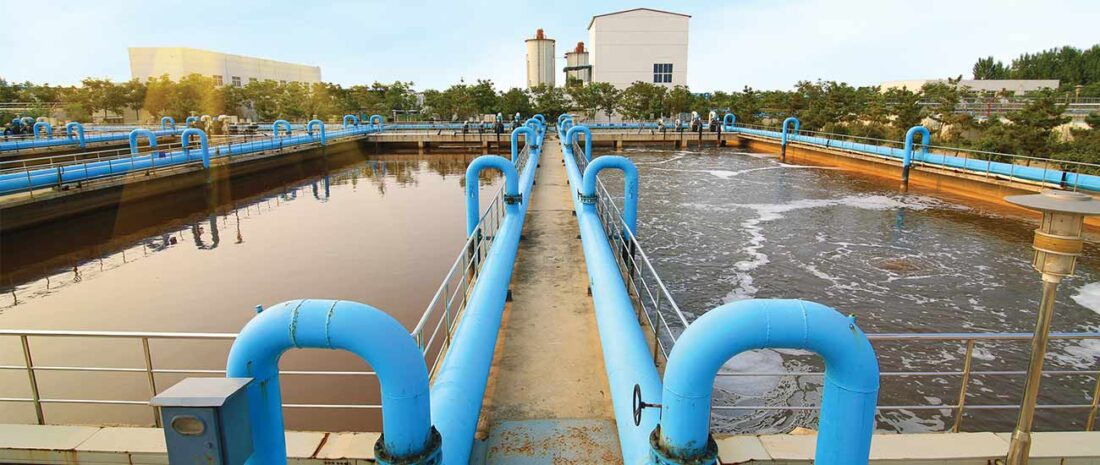
Ammonia removal is a key metric for assessing wastewater treatment facility performance. This is because ammonia contributes to aquatic life toxicity. Furthermore, nitrogen, along with phosphorus, is a driver of receiving water eutrophication. Eutrophication, an over-enrichment of nutrients, can be detrimental to environmental and public health. It can result in harmful algae blooms, dissolved oxygen depletion, fish kills, and other damaging impacts.
How does it work?
Ammonia can be biologically removed from wastewater by ammonia-oxidizing bacteria (AOB) and archaea (AOA) that convert ammonia to nitrite. This is the first step of nitrification; the second step involves the biological oxidation of nitrite to nitrate. Typical AOB and AOA are relatively slow-growing, aerobic, autotrophic organisms.
Traditional measurement tools
Ammonia oxidation is particularly susceptible to upsets resulting in the loss of nitrification. This can be caused by influent toxicity, seasonal changes such as decreased temperature and process changes such as decreased dissolved oxygen. Historically nitrification monitoring has been limited to measuring process parameters (DO, ORP, temperature, MLSS, SRT) and/or nitrogen species (ammonia, nitrite, nitrate, kjeldahl nitrogen). These all tend to be lagging indicators of nitrification loss.
Modern measurement tools
Modern microbiological tools allow operators and engineers to gain further insight into the process conditions that drive favorable process conditions for optimal nitrification rates.
Adenosine Triphosphate
Adenosine Triphosphate (ATP) can be used to measure the quantity of active biomass in a sample. It does not require culturing and therefore can be measured rapidly — less than 5 minutes — in the field or in a laboratory. Within a nitrification monitoring plan, ATP can be used to quantify bioreactor health and identify toxicity events.
qPCR
qPCR is a DNA-based tool that quantifies specific organisms or groups of organisms, such as AOA and AOB. qPCR results are generally expressed as cells/sample quantity (ie. mL), copies/ sample quantity or genomic units/ sample quantity. Traditionally this methodology was done by microbiologists in specialized labs, however recent developments have allowed this technology to be transferred to smaller, in-field units that can be used with pre-dispensed reagents with minimal experience in under 2 hours.
Next-Generation Sequencing
Next-Generation Sequencing (NGS) techniques, such as 16S rRNA sequencing, have been increasingly used in a wide range of applications. Within wastewater treatment facilities, NGS can be used to identify and quantify various beneficial and inhibitory organisms.
Understanding which organisms are present can help uncover the underlying ammonia-removal mechanisms (nitrification, comammox, anammox) and highlight the impact process changes have on community structure and function. For instance, different organisms may dominate the nitrifying community under different SRT. Understanding the relationship between community composition, SRT and nitrification rate can help to proactively optimize treatment operations. This method is still largely done within specialized labs; however, the price has decreased dramatically allowing it to be more economically accessible.
Learn more
These new tools are also challenging some of the preconceived notions of how treatment facilities should be operated to achieve nitrification. If you’re interested in learning more about LuminUltra’s suite of microbial monitoring tools and services, contact us today!