Why qPCR testing has emerged as a powerful alternative to culture testing
Two methods are widely used to test for the presence of Legionella bacteria in water supplies: culture testing and polymerase chain reaction (PCR) testing.
For over 150 years, culture testing has been used to diagnose disease. Along the way it’s become known as the “gold-standard” for detecting various strains of bacteria, including Legionella. But, over the last few decades, PCR (specifically, quantitative PCR or qPCR) testing has emerged as a powerful testing alternative for pathogen detection, preferred among many public health experts for its speed, accuracy and ease of use. A 2013 review found that, of 28 studies that used both methods, 26 detected Legionella at a higher rate using qPCR compared to culture testing.
Let’s dive into each of these methods and how they’re used in the detection of Legionella bacteria.
How is culture testing used to detect Legionella?
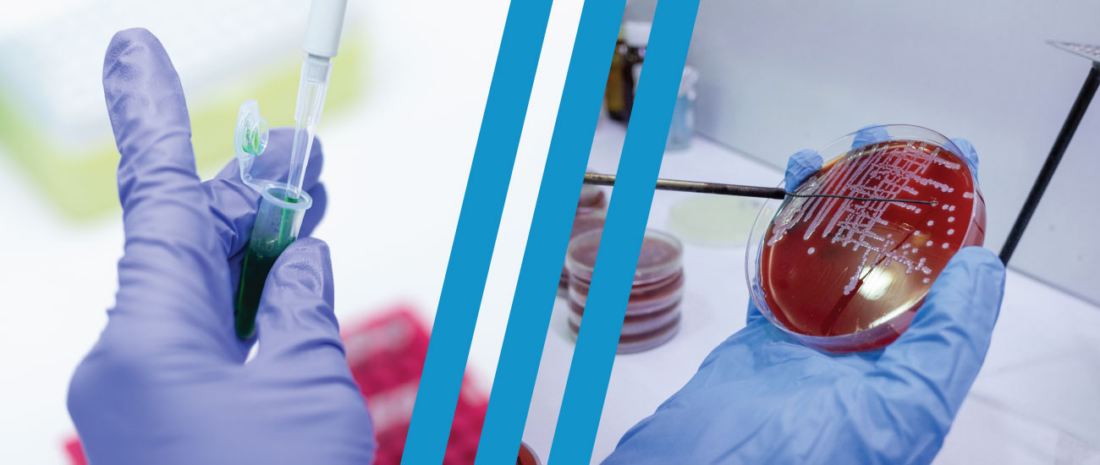
Culture testing is the traditional method for identifying the presence of Legionella bacteria in water sources, such as cooling towers. Culturing is the process of taking a sample of bacteria that’s present in a water source and allowing it to grow over a period of time in a petri dish, or another medium, until it’s visible and can be analyzed.
While culture testing is effective, it presents several drawbacks when used as the sole method to identify Legionella in a water source. The most significant of these is the time it takes (up to two weeks) to culture a sample. From the time an outbreak of Legionnaires’ disease is suspected, many more people can be exposed to the bacteria—shutting a building down during testing might not be a viable option—before its presence is confirmed by culture testing alone.
Another disadvantage of culturing is the specialized equipment and personnel required to do it properly. Legionella must be cultured in a highly-controlled environment—at the right temperature and provided with the right nutrients—to avoid multiplying the wrong microorganism, thus compromising the sample.
What’s more is that Legionella bacteria typically grow in a protective layer of larger cells, called a biofilm. The biofilm attaches itself to the inside walls or pipes and collects nutrients that are absorbed by the resident bacteria. When placed on a petri dish, the biofilm may prevent the growth of what’s inside it, which makes culturing them difficult.
How is qPCR testing used to detect Legionella?
Until recently, culture testing was considered to be the most reliable method of detecting Legionella in environments. But recent advancements in biotechnology have produced a method that’s been proven to provide faster and more accurate results: qPCR testing.
Rather than growing certain bacteria to observe them, qPCR testing isolates deoxyribonucleic acid (DNA) from a sample. This process requires a much smaller amount of the bacteria to be present, meaning the sample doesn’t need to be cultured to detect it. Because of this, a qPCR test can identify Legionella in a sample in under two hours — much faster than a culture test — while also detecting those bacteria that are viable but not culturable (VBNC), providing a more complete picture of risk.
Whether due to chemical treatments in water systems or sheltered in biofilms, bacteria can lie dormant and won’t grow in a petri dish. This presents problems for culture testing but, because DNA is still present whether bacteria are dormant or not, qPCR tests can identify dormant bacteria present in a sample.
The CDC recommends the use of both testing methods, where possible, in its guide for developing a response plan to Legionnaires’ disease.
How can I convert qPCR results to resemble culture methods?
Since many operators like to have results reported in a format that is familiar to them, it is possible to convert the gene copies/mL obtained from a qPCR reaction into an approximate cells/mL which is a calculation based on the average amount of gene copies per organism determined from literature investigations.
For LuminUltra qPCR assays like Legionella spp. or Total E. Coli which have specific genetically-defined targets, it is easy to convert from gene copies/mL to cells/mL because all the species covered are from the same genus, and are genetically similar.
Learn more about converting qPCR results.